
The Human Microbiome
New understanding of the complex human bacterial ecosystem has great implications for both oral and systemic health.
Introduction

In the 1990s, the possibility of a correlation between premature birth and periodontal diseases became a topic of research and discussion. Scientific study on this association also found support for relationships between periodontitis and other health problems. Today, the oral-systemic link is a well-known term. Research continues to delve deeper into the
cause of these associations—studying them at a molecular level. The examination of the microorganisms that reside on and within our bodies—the human microbiome—are vastly increasing our understanding of health and disease. In this month’s Sunstar Spotlight, Frances Doherty Genco, RDH, BS—a dental professional who contributes to this expanding knowledge base—shares her expertise on this fascinating topic.
Significant study has been dedicated to the molecular and genetic basis of the biology and individuality of humans. These advances have enabled scientists to investigate the microbes that inhabit the oceans, soil, human body, and much more. Through advanced DNA sequencing and powerful bioinformatics, science is now at the early stages of understanding the implications of the microbiome and its role in human survival. One day, these discoveries may lead to cures for diseases.
The study of the human microbiome provides a greater understanding of bacteria. This increased knowledge originated with the Human Genome Project, which began in 1990 and involved the collaboration of the National Institutes of Health (NIH), United States Department of Energy, and international partners. This concerted, public effort resulted in the sequencing of the entire human genome. Researchers found that humans have about 23,000 genes.1 This discovery led to the NIH?Human Microbiome Project, which strives to investigate microbial communities found at multiple human body sites for correlations between changes in the microbiome and human health.
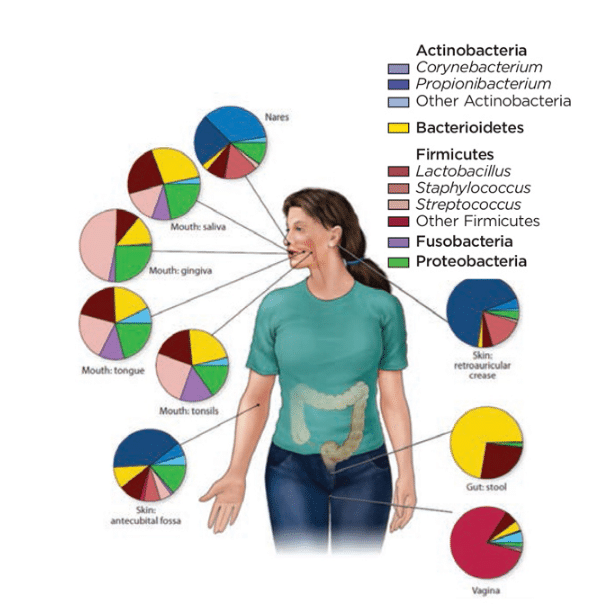
This research uses the same DNA sequencing methods to categorize bacteria in and on the human body and enables analysis of bacterial species and relative proportions without the need to culture.2 The key to success was to sequence the 16S ribosomal RNA gene, which enables identification of all bacteria at any site in the body. In 2012, the Human Microbiome Project reported the results of sequencing the bacteria of 242 healthy people at multiple sites, including the oral cavity, skin, gastrointestinal (GI) tract, nostrils, and vagina.3 The data showed that each site had a unique complex of hundreds of bacteria (Figure 1). The human microbiome contains about 100 trillion cells, which typically outnumber human cells by 10 to one. The human microbiome accounts for 99% of the genetic variation between individuals. The interaction of the microbiome with the human genome and environmental factors—such as diet, medication use, and exposure to toxins—holds great promise in addressing human disease.
The microbiome, especially the gut microbiome, has a huge impact on human survival—enabling the digestion of food, serving as the source of nutrients, and influencing development of the immune system. It affects mood and cognition, as well as diseases—including obesity, type 2 diabetes, Crohn’s disease, irritable bowel syndrome, and asthma. The microbiome may play a role in autoimmune diseases, such as type 1 diabetes, rheumatoid arthritis, muscular dystrophy, multiple sclerosis, fibromyalgia, as well as some cancers. Neurotransmitters known to act on the brain may be modified by the microbiome. By understanding these effects, improved treatment for schizophrenia, depression, bipolar disorder, autism, and other neurochemical imbalances may be possible. Dietary intake, circumstances of birth, and history of antibiotic usage affect the infant gut microbiome and upset the balance of bacteria that reside there.
The microbiome resists pathogens, but several pathogenic microbes can cause illness. The human microbiome consists primarily of commensal bacteria that call the human body home.3 The microbiome and the human host can be thought of as an ecosystem that, when healthy, is in balance. When the microbiome changes to a dysbiotic state, diseases such as dental caries, periodontal diseases, GI disorders, skin diseases, and neurological problems may occur. The dysbiotic state may harbor one or more of the bacterial pathogens that lead to diseases.
Understanding the microbiome may lead to new diagnostics, as well as preventive and treatment modalities for a variety of diseases. Determining the role of bacteria in maintaining health and causing disease is key to effectively treating bacterial infections. By grasping the functions and characteristics of the normal microbiome, novel strategies can be developed to modify the microbiome at specific sites with antibiotics, probiotics, prebiotics, or other therapies.
MICROBIOTA IN ORAL DISEASE
Studies of the microbiome in dental caries have revealed several important factors. For example, the microbiota in human dentinal caries in a Japanese population is complex, with three major clusters: a high Lactobacillus cluster, a mid-Lactobacillus cluster, and a low-Lactobacillus cluster.4 Using samples from the deepest area of each lesion, 11 genera were found to be major members of the total sequence. Prevotella was another prominent genus in dental caries.4 Another study demonstrated considerable heterogeneity in the microbiota associated with carious teeth.5 More studies are needed to better understand the microbiome in dental caries.
The role of subgingival microbiome in periodontal diseases has been studied extensively over the past several years. A series of previously unrecognized bacteria have been found in greater numbers among individuals with periodontal diseases. Many of the original organisms associated with periodontal diseases by culture methods have been confirmed by the studies of the subgingival microbiome.
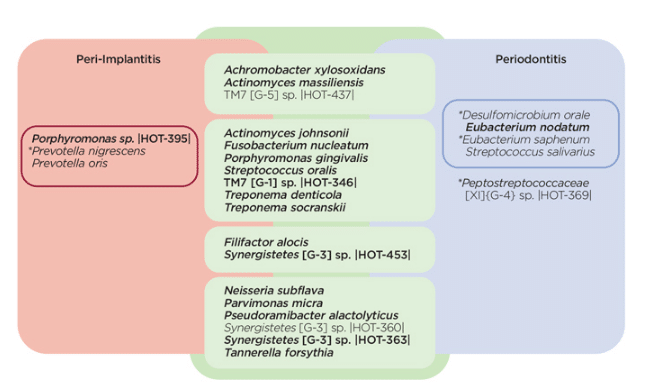
A recent study of the subgingival microbiota in patients with both periodontitis and peri-implantitis described the total microbiome of these conditions, in addition to the organisms that were unique (Figure 2).6 In this study of 20 patients, most of the predominant subgingival organisms are the same. However, some organisms are more prevalent in periodontitis lesions, including Peptostreptococcus and Desulfomicrobium orale. On the other hand, the proportion of P. nigrescens was higher in peri-implantitis lesions. Many of the organisms found in the diseased sites had been described previously by culture studies, including Fusobacterium nucleatum, Porphyromonas gingivalis, Treponema denticola, and Tannerella forsythia. Several organisms were newly associated with these diseases, including unnamed species designated as TM7, as well as Filifactor alocis and several Synergistes species. The role of these newly identified organisms remains to be determined. They may be causal or merely secondary invaders of the periodontal or peri-implantitis lesions.6
Most likely, several different clusters of subgingival pathogens are present in individual sites, and their elimination may require different therapies. Preventive and treatment procedures may also focus on restoring this microbiota using novel therapies, such as probiotic mixtures. Studies of the microbiome have suggested that the oral microbiota is stable over time and resists overgrowth of pathogens. Approaches to restoring the oral microbiome to health may prove critical in the prevention and treatment of periodontitis and peri-implantitis.
ORAL MICROBIOME AND SYSTEMIC DISEASES

Oral organisms have been detected in distant sites, such as the atheroma in patients with cardiovascular disease,7 colon tumors,8 and the placenta9 and fetal tissues of infants born prematurely or infants dying of fetal septicemia.10 These findings point to the possibility that bacteria are translocated from their original niche, such as the oral cavity, to distant sites in the body where they may contribute to disease processes. Future studies are needed to determine if this “mobile” microbiome can account for the associations of periodontal diseases with cardiovascular disease, adverse pregnancy outcomes, and other systemic conditions. The route by which bacteria from the periodontal subgingival microbiota translocates to other sites needs to be determined. The atheroma has been found to have a complex microbiome consisting of what appears to be organisms, possibly derived from the oral, gut, and skin microbiota (Figure 3).
Bacteria from the oral microbiota can travel to damaged heart tissue, leading to subacute bacterial endocarditis through short-lived bacteremias induced by dental procedures or even chewing—especially in patients with periodontitis. Recent evidence suggests that among patients with periodontitis, there is a chronic route of translocation. Phagocytic cells, such as dendritic cells, ingest but do not kill bacteria from the subgingival sites, enter the blood stream, and travel to the endothelium of the heart and contribute to atheroma formation.11
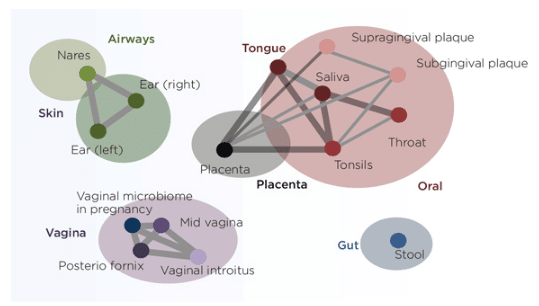
Similar mechanisms may explain the translocation of organisms to the placenta. A recent study12 found that a normal placenta harbors a complex microbiome, and the placental microbiota resembles the oral microbiota more than the microbiota from other sites on the body (Figure 4).
An intriguing association of F. nucleatum in colon tumors has been described.8 In vitro studies of F. nucleatum suggest that it can contribute to colon tumor formation.13 These findings need to be expanded to determine whether Fusobacterium are causal or secondary invaders of colon tumors. Also, future research is needed to determine the extent to which these organisms are truly derived from the oral cavity. These findings may well lead to an emphasis not only on reducing or eliminating pathogenic organisms from the subgingival area, but also on interfering with their route of translocation to distant diseased sites. This would represent a novel approach to preventing and treating conditions associated with periodontal diseases.
GUT MICROBIOME
The gut microbiome plays an important role in nutrition by degrading foods, and can be altered by antibiotics in early life—resulting in a dysbiotic gut microbiota that may contribute to allergies, asthma, obesity, and diabetes later in life.14
Food additives can alter the gut microbiome with deleterious effects. For example, artificial sweeteners have been shown to alter the gut microbiome, leading to obesity and prediabetes in a mouse model.15 Further studies are needed to support this finding.
CONCLUSION
It is the dawn of a new era in understanding the role of the human microbiome. Increased knowledge will likely lead to new diagnostic tests and novel preventive and treatment strategies, including attempts to restore the microbiome to its healthy state. This may include the use of prebiotics, foods that alter the microbiota; probiotics, live organisms that restore the normal microbiota; or symbiotics, which are a combination of the two. A new generation of targeted antibacterial agents to substitute for broad spectrum antibiotics are actively being investigated. Several are being clinically tested for diseases of the intestine, skin, and oral cavity. Improved understanding of this complex ecosystem holds great promise in managing oral health problems and the several serious chronic conditions linked to oral diseases.
ACKNOWLEDGEMENT
The author would like to thank Robert J. Genco, DDS, PhD, for his assistance in the preparation of this manuscript.
REFERENCES
- National Human Genome Research Institute. An Overview of the Human Genome Project. Available at: genome.gov/12011238. Accessed December 12, 2014.
- Cho I, Blaser MJ. The human microbiome: at the interface of health and disease. Nat Rev Genet. 2012;13:260–270.
- Grice EA, Segre JA. The human microbiome: our second genome. Annu Rev Genomics Hum Genet. 2012;13:151–170.
- Obata J, Takeshita T, Shibata Y, et al. Identification of the microbiota in carious dentin lesions using 16S rRNA gene sequencing. PLoS One. 2014;9:e103712.
- Peterson SN, Snesrud E, Liu J, et al. The dental plaque microbiome in health and disease. PLoS One. 2013;8:e58487.
- Maruyama N, Maruyama F, Takeuchi Y, Aikawa C, Izumi Y, Nakagawa I. Intraindividual variation in core microbiota in peri-implantitis and periodontitis. Sci Rep. 2014;4:6602.
- Armingohar Z, Jørgensen JJ, Kristoffersen AK, Abesha-Belay E, Olsen I. Bacteria and bacterial DNA in atherosclerotic plaque and aneurysmal wall biopsies from patients with and without periodontitis. J Oral Microbiol. 2014;15:6.
- Castellarin M, Warren RL, Freeman JD, et al. Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res. 2012;22:299–306.
- Fardini Y, Chung P, Dumm R, Joshi N, Han YW. Transmission of diverse oral bacteria to murine placenta: evidence for the oral microbiome as a potential source of intrauterine infection. Infect Immun. 2010;78:1789–1796.
- Han YW, Wang X. Mobile microbiome: oral bacteria in extra-oral infections and inflammation. J Dent Res. 2013;92:485–491.
- Carrion J, Scisci E, Miles B, et al. Microbial carriage state of peripheral blood dendritic cells (DCs) in chronic periodontitis influences DC differentiation, atherogenic potential. J Immunol. 2012;189:3178–3187.
- Aagaard K, Ma J, Antony KM, Ganu R, Petrosino J, Versalovic J. The placenta harbors a unique microbiome. Sci Transl Med. 2014;6:237ra65.
- Rubinstein MR, Wang X, Liu W, Hao Y, Cai G, Han YW. Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-Cadherin/β-Catenin signaling via its FadA adhesion. Cell Host Microbe. 2013;14:195–206.
- Cox LM, Yamanishi S, Sohn J, et al. Altering the intestinal microbiota during a critical development window has lasting metabolic consequences. Cell. 2014;158:705–721.
- Suez J, Korem T, Zeevi D, et al. Artificial sweeteners induce glucose intolerance by altering the gut microbiota. Nature. 2014;514:181–186.
From Dimensions of Dental Hygiene. January 2015;13(1):39–42.

